Home
My projects
Sometimes I create stuff
SimMod
Detection of Composite Communities in Multiplex Biological Networks
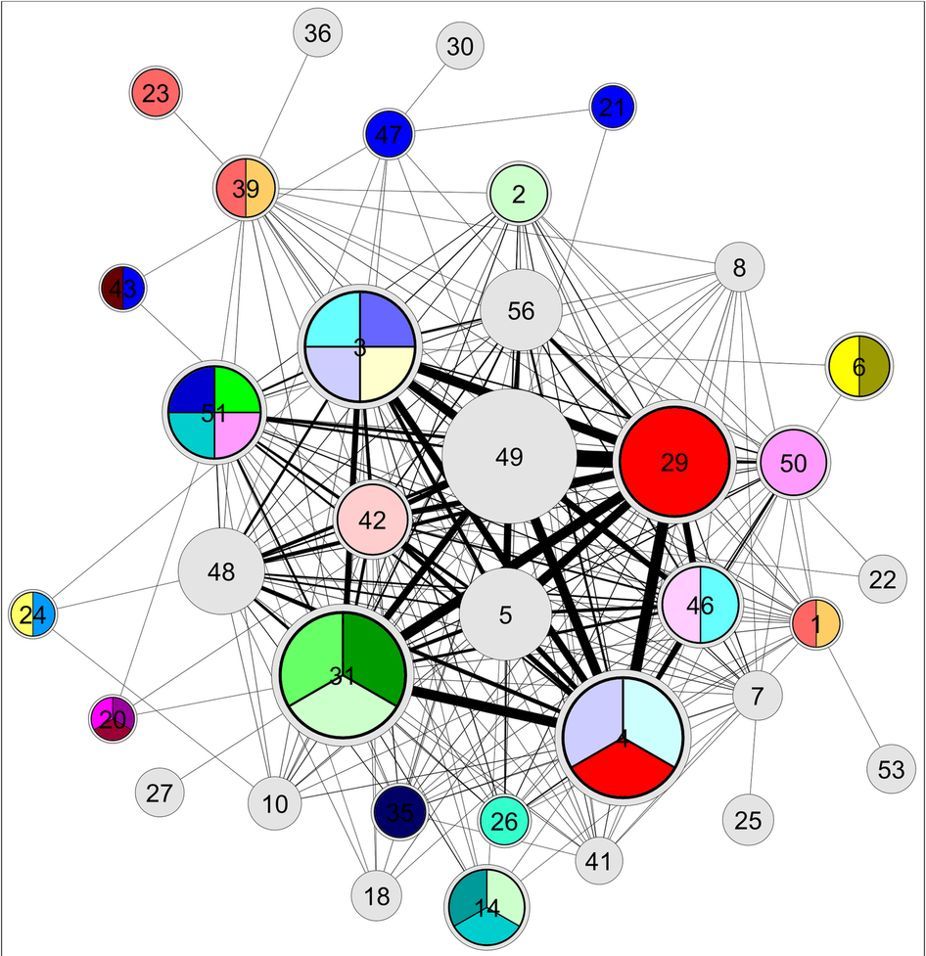
 Cellular organisation is assumed to be modular, with each module driving a distinct biological process. The detection of community structure is a widely accepted means of investigating the principles governing biological systems. Recent efforts are exploring ways in which multiple data sources can be integrated to generate a more comprehensive model of cellular interactions, leading to the detection of more biologically relevant communities. Communities, also known as modules, have been shown to comprise groups of biomolecules that physically interact, are functionally cohesive, co-regulated or correspond to biological pathways. Community detection applications have linked molecular compounds with disease, correlated the organisation of cancer signalling networks with patient survival rate6 and identified functional modules related to coronary artery disease.
Cellular organisation is assumed to be modular, with each module driving a distinct biological process. The detection of community structure is a widely accepted means of investigating the principles governing biological systems. Recent efforts are exploring ways in which multiple data sources can be integrated to generate a more comprehensive model of cellular interactions, leading to the detection of more biologically relevant communities. Communities, also known as modules, have been shown to comprise groups of biomolecules that physically interact, are functionally cohesive, co-regulated or correspond to biological pathways. Community detection applications have linked molecular compounds with disease, correlated the organisation of cancer signalling networks with patient survival rate6 and identified functional modules related to coronary artery disease.
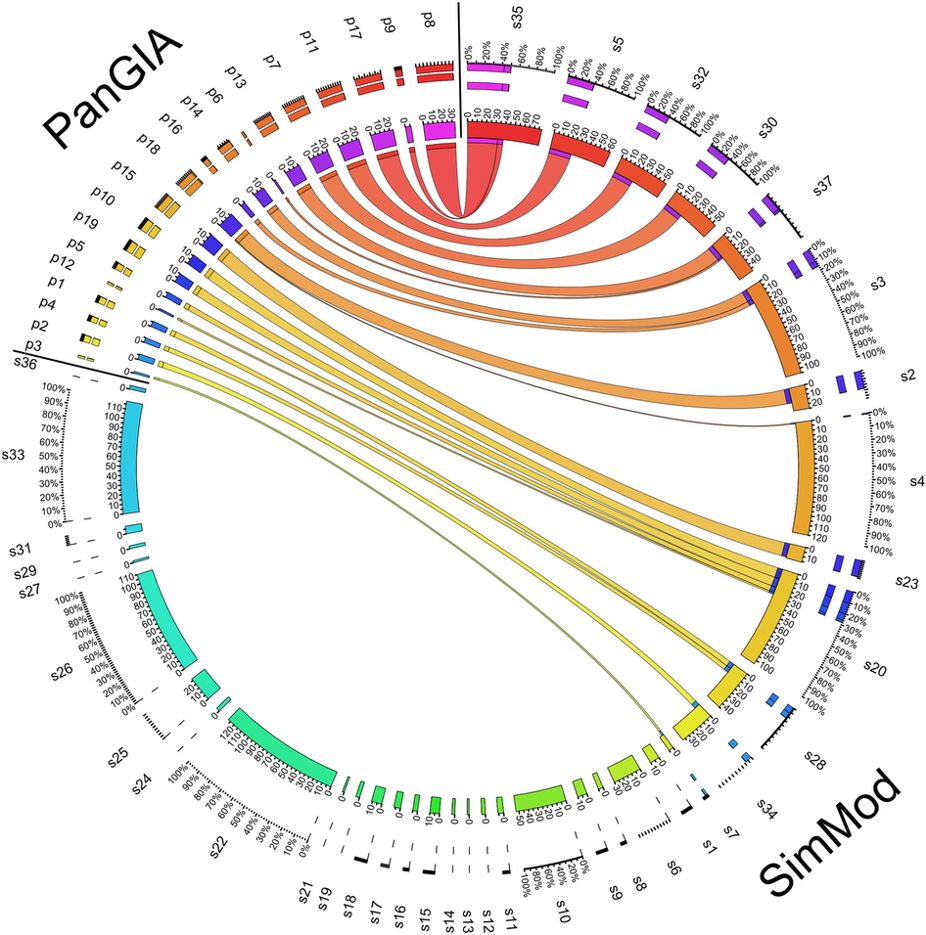
 In this work, we propose a mathematical programming model to cluster multiplex biological networks, i.e. multiple network slices, each with a different interaction type, to determine a single representative partition of composite communities. Our method, known as SimMod, is evaluated through its application to yeast networks of physical, genetic and co-expression interactions. A comparative analysis involving partitions of the individual networks, partitions of aggregated networks and partitions generated by similar methods from the literature highlights the ability of SimMod to identify functionally enriched modules. Functional enrichment analysis (Gene Ontology enrichment) has been employed to assess the resulting modules and we find that SimMod discovers relevant biologically functional modules with fairly good enrichment scores. It is further shown that SimMod offers enhanced results when compared to existing approaches without the need to train on known cellular interactions.
In this work, we propose a mathematical programming model to cluster multiplex biological networks, i.e. multiple network slices, each with a different interaction type, to determine a single representative partition of composite communities. Our method, known as SimMod, is evaluated through its application to yeast networks of physical, genetic and co-expression interactions. A comparative analysis involving partitions of the individual networks, partitions of aggregated networks and partitions generated by similar methods from the literature highlights the ability of SimMod to identify functionally enriched modules. Functional enrichment analysis (Gene Ontology enrichment) has been employed to assess the resulting modules and we find that SimMod discovers relevant biologically functional modules with fairly good enrichment scores. It is further shown that SimMod offers enhanced results when compared to existing approaches without the need to train on known cellular interactions.
Check out the publication in Nature - Scientific Reports.