Home
My projects
Sometimes I create stuff
HyperDC
Pathway-level disease data mining through hyper-box principles
In microarray data analysis, traditional methods that focus on single genes are increasingly replaced by methods that analyse functional units corresponding to biochemical pathways, as these are considered to offer more insight into gene expression and disease associations. However, the development of robust pipelines to relate genotypic functional modules to disease phenotypes through known molecular interactions is still at its early stages.
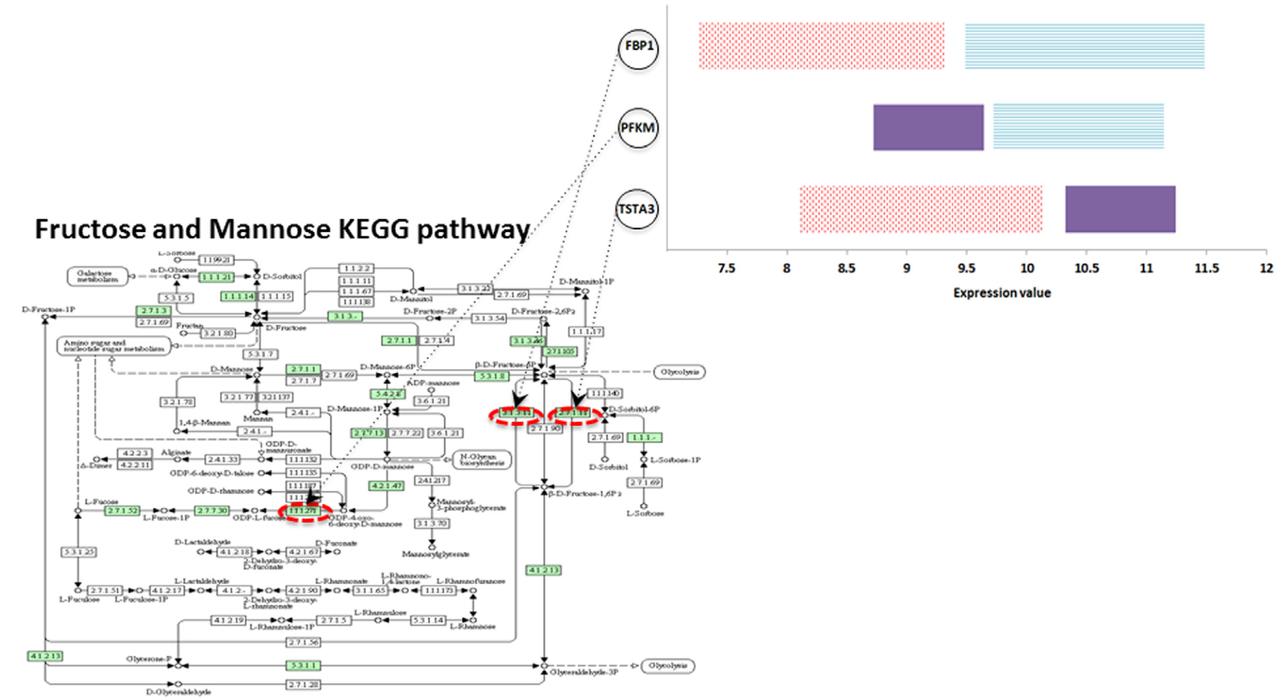 In this article we first discuss methodologies that employ groups of genes in disease classification tasks that aim to link gene expression patterns with disease outcome. Then we present a pathway-based approach for disease classification through a mathematical programming model based on hyper-box principles. Association rules derived from the model are extracted and discussed with respect to pathway-specific molecular patterns related to the disease. Overall, we argue that the use of gene sets corresponding to disease-relevant pathways is a promising route to uncover expression-to-phenotype relations in disease classification and we illustrate the potential of hyper-box classification in assessing the predictive power of functional pathways and uncover the effect of specific genes in the prediction of disease phenotypes.
In this article we first discuss methodologies that employ groups of genes in disease classification tasks that aim to link gene expression patterns with disease outcome. Then we present a pathway-based approach for disease classification through a mathematical programming model based on hyper-box principles. Association rules derived from the model are extracted and discussed with respect to pathway-specific molecular patterns related to the disease. Overall, we argue that the use of gene sets corresponding to disease-relevant pathways is a promising route to uncover expression-to-phenotype relations in disease classification and we illustrate the potential of hyper-box classification in assessing the predictive power of functional pathways and uncover the effect of specific genes in the prediction of disease phenotypes.
- Mathematical programming method based on hyper-box principles is applied in pathway-level disease classification.
- The model entails advantages of descriptive power and flexibility and can extract multiple phenotype-responsive genes.
- Prediction accuracy on breast cancer and psoriasis gene expression is completitive against other classification approaches.
- The use of pathway gene sets improves classification accuracy, as compared against randomly selected gene groups.
Check it out in Mathematical Biosciences here.